November 15, 2011 (Vol. 31, No. 20)
URL:
http://t.caspur.it/ASPicDB/
Rating:
Strong Points: Links page to similar resources, simple layout
Weak Points: Nothing major
Summary:
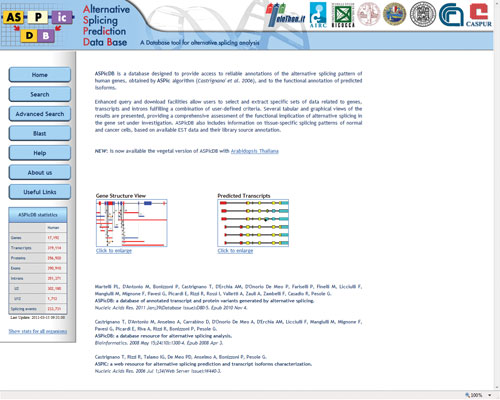
During processing, mRNA transcripts may be spliced to yield multiple final products; this phenomenon is known (appropriately enough) as alternative splicing. The tricky thing is, one cannot know how a transcript will be spliced simply by looking at the gene sequence…unless, of course, you have a handy, dandy alternative splicing fortune teller like ASPicDB (more formally, the Alternative Splicing Prediction Data Base). For each query, ASPicDB returns a gene structure view, predicted transcripts, a transcript table, predicted proteins, a protein table, predicted splice sites, and an intron table. There is also limited data regarding gene expression. If one alternative splicing prediction algorithm isn’t satisfying enough, one can check out (dare I say, alternative) splicing databases under the “useful links” tab. I predict you’ll find this site quite useful…